Quick usage¶
Running GAUDI jobs is quite easy with gaudi.cli.gaudi_run:
gaudi run /path/to/some_file.gaudi-input
To learn how to create input files, go to Input files, and make sure to check the tutorials!
After the job is completed, you can check the results with gaudi.cli.gaudi_view:
gaudi view /path/to/some_file.gaudi-output
You can choose between two molecular visualization tools: Chimera itself (using GaudiView extension), or our in-house GUI, GAUDInspect.
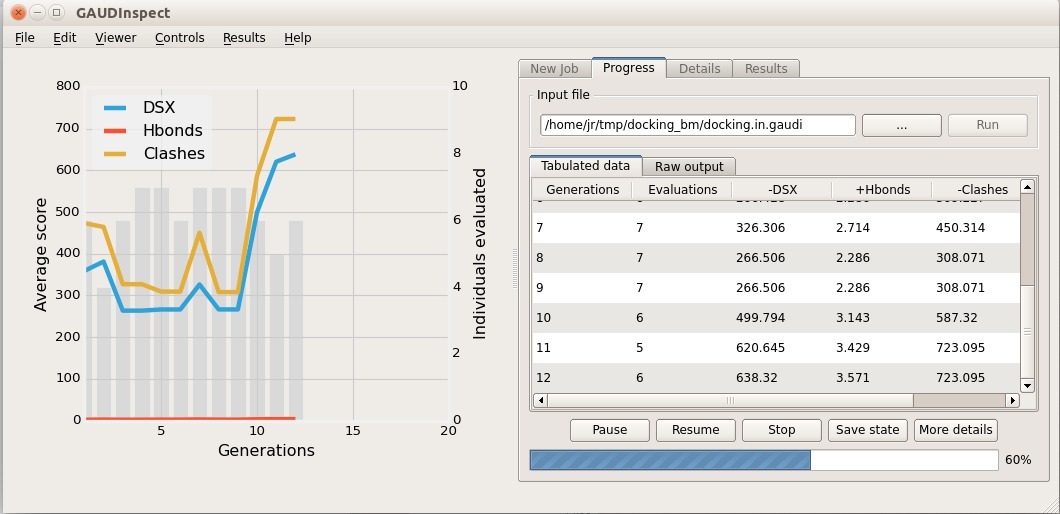
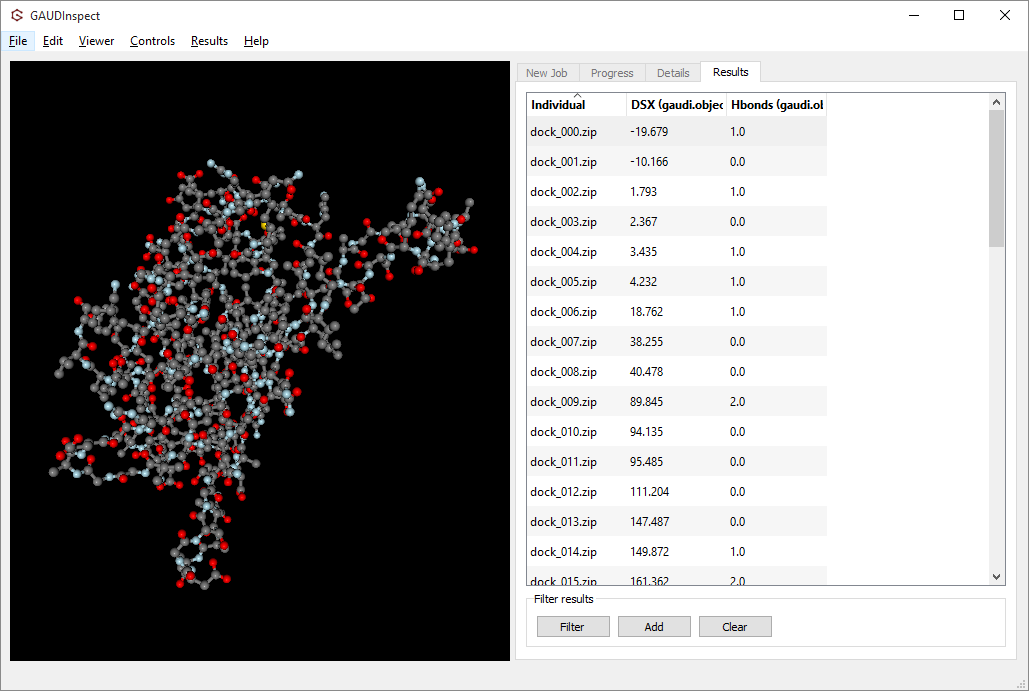
Both tools support lazy-load, filtering and sorting, so choose whichever you prefer. A quick tutorial on GaudiView is available here: